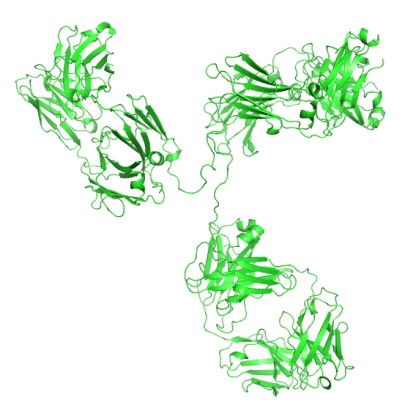
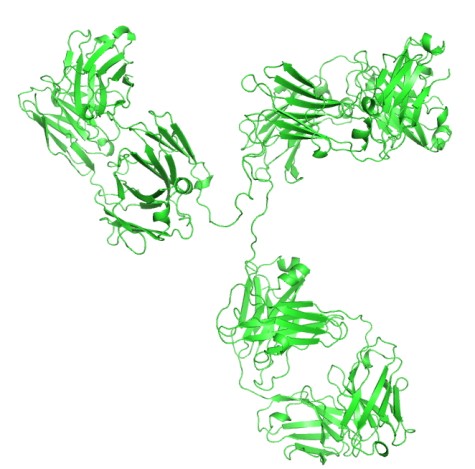
Antibodies can be dissected into smaller antigen binding fragments, by proteolysis or genetic engineering, to produce mono or multivalent fragments. These are mainly based on either Fab fragments or the single chain Fv (scFv) as building blocks. The Fv portion of an IgG, consisting of the VH and VL domains, is the smallest fragment that maintains the full binding capacity of the intact antibody.
We have established a toolbox of methods to transfer the genetic information for antibodies from B-cells of animals to yeast. Animals are immunized with a target protein of interest and the mRNA is extracted from B-cells, converted into DNA, amplified and transferred into a yeast background. This results in a set of yeast cells, each displaying an antibody variant in multiple copies on the cell surface. High throughput fluorescence-activated cell sorting (FACS) is used to identify antibodies with prescribed binding characteristics.
Nowadays, 5% of all newly diagnosed malignancies are lymphoma related. Current immunotherapies rely on targeting tumor markers over-expressed on lymphoma cells, but also present on healthy cells. As a consequence, standard treatments often lead to severe side effects. Therefore, precision medicine based on a personalized therapy might be beneficial for lymphoma treatment. In B-cell lymphomas, the cancer cells express a tumor-specific and functionally active B-cell receptor (BCR), also referred to as idiotype (Id), which differs from the homologue structure of healthy cells. In contrast to conventional antibodies from mammals, cartilaginous fish have a unique form of heavy chain-only antibodies called immunoglobulin new antigen receptor (IgNAR), with a variable domain (vNAR) responsible for antigen binding. Binders specifically directed towards the BCR of different lymphoma B-cell lines have been identified from yeast-displayed, semi-synthetic vNAR libraries. In ongoing work, we are focused on the design of several anti-idiotypic vNAR formats for the development of a personalized lymphoma treatment in a cost and time efficient manner, while drastically reducing side effects.
In addition to antibodies with the classical composition of the heavy and the light chains, the adaptive immune repertoire of sharks also includes a heavy-chain only isotype, where antigen binding is mediated exclusively by a small and highly stable domain, referred to as vNAR. Due to their high affinity and specificity combined with their small size, high physicochemical stability and low-cost production, vNAR fragments have evolved in recent years, as promising target-binding scaffolds that can be tailor-made for applications in medicine and biotechnology.
High-affinity shark-derived nanobodies (vNARs) have been identified for a variety of different antigens. By using semi-synthetic libraries, we were able to implement special features such as pH-responsive binding. vNARs are particularly useful for the generation of anti-idiotypic binders directed against therapeutic antibodies. We developed a novel route for broadening the therapeutic window of cancer antibodies by masking their antigen-binding site with an anti-idiotypic vNAR. The ultimate goal is to generate a vNAR-antibody fusion protein that renders the antibody inactive in normal tissue while being activated in the tumor microenvironment.
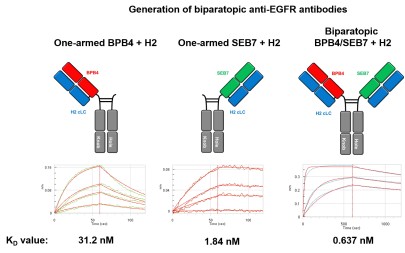
Recently, chicken-derived antibodies became interesting therapeutic and diagnostic entities due to their broad epitope coverage and the simple library generation. The goal of our research is the development of next-generation antibodies, like bispecific and biparatopic mAbs. To facilitate the correct chain pairing, the Knob-into-Hole technology as well as common light chains (cLC) are utilized. Light chain shuffling to improve affinity and library-based humanization approaches are used to generate antibody candidates with tailor-made properties. Furthermore, library-based methods for the generation of pH-responsive bispecific antibodies are explored that can facilitate new modes of action. The establishment of a novel yeast bio-panning-based, FACS-assisted screening method expanded the repertoire of screening technologies for cLC-based antibodies.
|
Facile generation of antibody heavy and light chain diversities for yeast surface display by Golden Gate Cloning. Roth L, Grzeschik J, Hinz SC, Becker S, Toleikis L, Busch M, Kolmar H, Krah S, Zielonka S. Biol Chem. 2019 Feb 25;400(3):383-393. doi: 10.1515/hsz-2018-0347. |
|
Yeast Surface Display in Combination with Fluorescence-activated Cell Sorting Enables the Rapid Isolation of Antibody Fragments Derived from Immunized Chickens. Grzeschik J, Yanakieva D, Roth L, Krah S, Hinz SC, Elter A, Zollmann T, Schwall G, Zielonka S, Kolmar H. Biotechnol J. 2019 Apr;14(4):e1800466. doi: 10.1002/biot.201800466. |
|
A Streamlined Approach for the Construction of Large Yeast Surface Display Fab Antibody Libraries. Krah S, Grzeschik J, Rosowski S, Gaa R, Willenbuecher I, Demir D, Toleikis L, Kolmar H, Becker S, Zielonka S. Methods Mol Biol. 2018;1827:145-161. doi: 10.1007/978-1-4939-8648-4_8. |
|
Construction of Histidine-Enriched Shark IgNAR Variable Domain Antibody Libraries for the Isolation of pH-Sensitive vNAR Fragments. Könning D, Hinz S, Grzeschik J, Schröter C, Krah S, Zielonka S, Kolmar H. Methods Mol Biol. 2018;1827:109-127. doi: 10.1007/978-1-4939-8648-4_6. |
|
Semi-synthetic vNAR libraries screened against therapeutic antibodies primarily deliver anti-idiotypic binders. Könning D, Rhiel L, Empting M, Grzeschik J, Sellmann C, Schröter C, et al. Scientific reports. 2017;7(1):9676. |
Sactipeptides belong to the class of ribosomal synthesized and posttranslational modified peptides (RiPPs) of bacterial origin. These unique peptides contain at least one thioether bridge, which is a linkage between the thiol of a cysteine residue and the α-carbon of a partner amino acid. This unique linkage provides these peptides with extraordinary chemical stability and conformational rigidity. The stability and the antimicrobial activity of sactipeptides (also known as sactibiotics) makes them highly interesting research targets. Since only few of these peptides could be identified up to date, our research focuses on the identification and structural and functional characterization of novel sactipeptides. Therefore, different predicted sactipeptide gene clusters, which were found via genome mining, are analyzed for the presence of the oxygen sensitive modifying radical SAM enzymes and sactipeptides.
For further reading see our Angewandte Chemie paper
Furthermore, we are working on the establishment of a high throughput platform for the screening of sactipeptide variants with novel functionalities. Recently, Urban et al., 2017 and Hetrick et al., 2018 developed a lanthipeptide screening platform employing phage display and yeast display, respectively. However, a high throughput screening of sactipeptides has yet to be achieved.
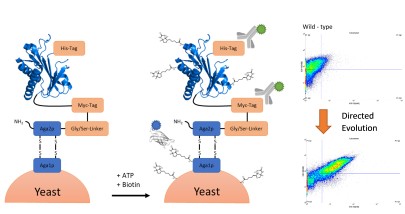
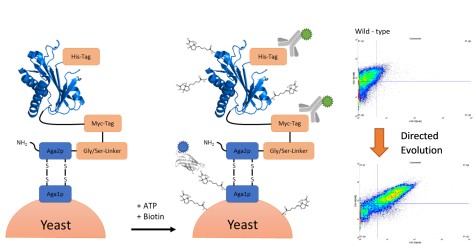
Protein-protein interactions studies enable a better understanding of processes happening in whole cells. BioID-screen (proximity-dependent biotin identification) is useful tool to achieve this purpose. Fusing a promiscuous biotin ligase to a protein-of-interest (POI) leads to biotinylation of all proteins interacting with it. The introduced biotin enables easy purification and identification of the interaction partners. Up to date, biotin ligases used have either a large molecular weight or a slow reaction rate. To solve this problem, we performed a directed evolution of a small biotin ligase derived from A. aeolicus in order to improve catalytic properties. After the generation of an Error-Prone PCR based yeast-surface display library, we identified two mutants exhibiting higher turnover than any other ligase known.
For details see our recent publication.
We are looking for methods to preserve the native heavy chain / light chain pairing of B cells during generation of yeast surface display (YSD) libraries. To this end, single B cells are encapsulated with poly(dT) magnetic beads in lysis buffer using water-in-oil emulsion microfluidics. After cell lysis, the beads carrying mRNA of single cells are isolated and resuspended in reverse transcription PCR (RT-PCR) buffer and re-emulsified for PCR. Amplicons for VH and VL domains are covalently connected either via SOE-PCR or ligase chain reaction (LCR). The final PCR products can then be assembled into bidirectional promoter vectors for YSD of Fab fragments via Golden Gate Assembly.
Early stage functional screening of biologics is beneficial for the discovery and generation of new potent first-in class drugs. Classical in vitro microtiter plate-based functional assays, however, require reformatting of the test molecules, heterologous expression and purification, limiting the throughput and extending project time lines. We investigate the establishment of ultra-high-throughput functional screen by compartmentalization of mammalian reporter cells together with S. cerevisiae secreting a library of test molecules, in millions of agarose-containing microdroplets. Utilizing droplet-based microfluidics, co-encapsulation in mono-dispersed and highly homogeneous droplets is achieved. Co-cultivation of the encapsulated cells in water-in-oil emulsion leads to accumulation of the secreted test molecules from the yeast within the droplet, enabling specific activation of the reporter cells, thus ensuring genotype-phenotype coupling. Following agarose solidification, activated reporter cells and the corresponding yeast cells are sorted in an ultra-high-throughput manner utilizing a commercially available FACS, without need of extensive optimization and adaptation. With this project, are aiming to develop a system for early-stage discovery of agonists, antagonists, or even cytosol-internalizing antibodies.
|
Genome mining for ribosomally synthesized and post-translationally modified peptides (RiPPs) in anaerobic bacteria. Letzel AC, Pidot SJ, Hertweck C. BMC Genomics 2014,15:983 |
|
An improved smaller biotin ligase for BioID proximity labeling. Kim DI, Jensen SC, Noble KA, Kc B, Roux KH, Motamedchaboki K, Roux KJ. Mol Biol Cell. 2016 Apr 15;27(8):1188-96. |
|
Ultra-high-throughput sequencing of the immune receptor repertoire from millions of lymphocytes. McDaniel JR, DeKosky BJ, Tanno H, Ellington AD, Georgiou G. Nature Protocols, 11, 429–442(2016) |
|
A One Pot, One Step, Precision Cloning Method with High Throughput Capability. Engler C, Kandzia R, Marillonnet S. PLos ONE, 2008, 3, 11, e364. |
Up to date, the repertoire of inducible transcription activators in eukaryotic cells is limited and moreover, they possess some disadvantages. While Tet-Off/On based systems can be regulated tightly, they still require clinically relevant antibiotics. For surface display of proteins on yeast cells (YSD), induction via galactose achieves strong transcription activation, but limits cell growth due to the required absence of glucose. Thus, research of metabolic pathways and the construction of complex genetic circuits would benefit from the design of novel transcription regulators, individually switchable with less demanding inducers and implementable in desired numbers per cell. The 2017 published “Arbitrium” system of bacteriophages could provide such proteins.
|
A tightly regulated and adjustable CRISPR-dCas9 based AND gate in yeast. Hofmann A, Falk J, Prangemeier T, Happel D, Köber A, Christmann A, Koeppl H, Kolmar H. Nucleic Acids Research, 47, 1, 2019, 509–520 |
|
Communication between viruses guides lysis–lysogeny decisions. Erez Z, Steinberger-Levy I, Shamir M, Doron S, Stokar-Avihail A, Peleg Y, Melamed S, Leavitt A, Savidor A, Albeck S, Amitai G, Sorek R, Nature, 2017, 541, 488–493 |
Covalent linkage of certain peptidic units, being an alternative or a complementary strategy to recombinant production, often becomes a method of choice for the synthesis of sophisticated macromolecular constructs with tailored properties. To date a vast number of bioconjugates and the respective techniques have been reported, ranging from relatively simple fluorescently or small-molecule-labelled peptides to complex, multifunctional architectures like antibody-drug conjugates.
Antibody-drug conjugates (ADCs) are an emerging class of molecules for the targeted treatment of cancer. They combine the remarkable specificity of therapeutic antibodies with the high potency of small molecular drugs by covalent linkage of both molecules to overcome the harsh side-effect of conventional chemotherapy. However, current methodologies for the coupling of the cytotoxic warhead to the tumor-targeting antibody suffer from off-target conjugation and low specificity. In this research, an enzymatic strategy employing microbial transglutaminase from Streptomyces mobaraensis that is capable of forming highly stable isopeptide bonds between the side-chains of glutamine and lysine residues, is used to circumvent these limitations.
The equipment of antibodies, nanobodies or other molecules applied in targeted therapy with the hydrophilic polysaccharide dextran allows for higher loading of those targeting molecules with cargoes of interest (Schneider et al., 2019). Orthogonal coupling systems, especially click reactions, enable the site specific introduction of desired active substances to the dextran scaffold. The subsequent attachment of dextran to the targeting anti- or nanobody can be carried out either via bioconjugation utilizing enzymes like microbial transglutaminase (mTG), sortase A (Sort A), or by click chemistry.
These conjugates feature high drug payload whilst maintaining or even enhancing hydrophilicity of the immunoglobulin towards the preservation of pharmacokinetics. Therefore, dextran functionalization of already approved targeting molecules is a promising strategy to engineer novel and highly efficient anti-cancer agents.
The immobilization of sensing molecules, e.g. enzymes or antibodies, on the surface of lignocellulosic fibers is crucial for any paper-based bioanalytical application. Up to date, it is not trivial to immobilize biomolecules in a directed, orthogonal fashion on the surface of paper fibers with a high loading density by common chemical conjugation strategies.
We investigate a general method for such a directed conjugation of biomacromolecules to paper fibers, based on a chemical pre-functionalization of the fibers with thermally stable peptides as selective linker-molecules. After papermaking, site-specific immobilization of bioactive proteins will be performed by a mild enzyme-catalyzed bioconjugation. The results of this work will enable future research towards detailed understanding of microfluidic papers with tailored biocatalytic properties.
|
Recent progress in transglutaminase-mediated assembly of antibody-drug conjugates. Schneider H, Deweid L, Avrutina O, Kolmar H. Anal Biochem. 2020, 15;595:113615. |
|
Directed Evolution of a Bond-Forming Enzyme: Ultrahigh-Throughput Screening of Microbial Transglutaminase Using Yeast Surface Display. Deweid L, Neureiter L, Englert S, Schneider H, Deweid J, Yanakieva D, Sturm J, Bitsch S, Christmann A, Avrutina O, Fuchsbauer HL, Kolmar H. Chemistry. 2018,12, 24(57):15195-15200. |
|
Light‐Controlled Chemoenzymatic Immobilization of Proteins towards Engineering of Bioactive Papers. Valentina Hilberg V, Avrutina O, Ebenig A, Desislava Yanakieva D, Meckel T, Biesalski M, Kolmar H. Chem. Eur. J., 2019, 25,1746 –1751. |
|
A Chemoenzymatic Approach to Protein Immobilization onto Crystalline Cellulose Nanoscaffolds. Uth C, Zielonka S, Hörner S, Rasche N, Plog A, Orelma H, Avrutina O, Zhang K, Kolmar H. Angew. Chem. Int. Ed., 2014, 53, 12618 –12623. |
The therapeutic potential of biomacromolecules (e.g. proteins, nucleic acids) is limited, due to their inability to reach intracellular targets. Hence, increasing effort is being directed at the development of molecular transporters, like cell-penetrating peptides (CPPs) or liposomes to overcome this handicap. Despite the rapid advance in this field, many transporters still suffer from shortcomings like endosomal entrapment and cytotoxicity. We are focusing on the delivery of peptide nucleic acids (PNA), a synthetic DNA analogue, and proteins primarily to eukaryotic cells using CPPs. Additionally, we are investigating whether multiple covalent attachment of CPPs on the polysaccharide dextran scaffold results in higher efficacy.
|
Recent Advances in Cell Penetrating Peptide-Based Anticancer Therapies. Habault J, Jean-Luc Poyet JL Molecules 2019, 24(5), 927 |
|
Soy Lecithin-Derived Liposomal Delivery Systems: Surface Modification and Current Applications. Ngoc Thuy Trang Le, Van Du Cao, Thi Nhu Quynh Nguyen, Thi Thu Hong Le, Thach Thao Tran, Thai Thanh Hoang Thi Int. J. Mol. Sci. 2019, 20(19), 4706 |
|
Peptide Nucleic Acids (PNA): Synthesis, properties and potential applications. Hyrup B, Nielsen PE Bioorganic & Medicinal Chemistry, 4, 1, 1996, 5-23 |
|
TRAIL‐Inspired Multivalent Dextran Conjugates Efficiently Induce Apoptosis upon DR5 Receptor Clustering. Hendrik Schneider H, Yanakieva D, Macarrón A, Deweid L, Becker B, Englert S, Avrutina O, Kolmar H ChemBioChem, 2019, 20, 3006 –3012 |
|
Multivalent dextran hybrids for efficient cytosolic delivery of biomolecular cargoes. Becker B, Englert S, Schneider H, Yanakieva D, Hofmann S, Dombrowsky C, Macarrón Palacios A, Bitsch S, Elter A, Meckel T, Kugler B, Schirmacher A, Avrutina O, Diederichsen U, Kolmar H. J Pept Sci. 2021 Apr;27(4):e3298. doi: 10.1002/psc.3298. Epub 2021 Jan 17. |
In collaboration with the pharma company Fuse Biotherapeutics we are developing tailor-made monoclonal antibodies for therapeutic use with focus on applications in oncology.
Antibodies are a successful class of therapeutics. Over 94 are currently approved, more than 270 in clinical trials.
We are using state-of-the-art technologies for antibody isolation and functional validation. Antibody candidates with optimal target binding are further characterized for their ability to be effective in various cell-based and animal disease models.
Sulfotools GmbH, a spin-off of TU Darmstadt, has developed a novel technology for the sustainable and cost-effective production of peptides. The Clean Peptide Technology (CPT) is based on novel highly water soluble protecting groups, the use of which in peptide synthesis, allows for the substitution of so far required hazardous organic solvents with water.
Merck Healthcare KGaA
Since more than ten years we have an extremely fruitful collaboration with the Merck Healthcare in Darmstadt. More than 65 publications are indicative of productive common research activitites.